Transfected Stable Cell Lines
Reliable | High-Performance | Wide Rage
Precision reporter, kinase, immune receptor, biosimilar, Cas9, and knockout stable cell lines for diverse applications.
Transfected Stable Cell Lines
Reliable | High-Performance | Wide Rage
Precision reporter, kinase, immune receptor, biosimilar, Cas9, and knockout stable cell lines for diverse applications.
Premade Virus Particles
Ready-to-Use | High Titer | Versatile Applications
Premade AAV, adenovirus, lentivirus particles, safe, stable, in stock.
Virus-Like Particles (VLPs)
Stable | Scalable | Customizable
Advanced VLPs for vaccine development (Chikungunya, Dengue, SARS-CoV-2), gene therapy (AAV1 & AAV9), and drug screening (SSTR2, CCR5).
Oligonucleotide Products
Precise | High Yield | Tailored Solutions
Accelerate your research with cost-effective LncRNA qPCR Array Technology.
RNA Interference Products
Targeted | Potent | High Specificity
Human Druggable Genome siRNA Library enables efficient drug target screening.
Recombinant Drug Target Proteins
Authentic | Versatile | Accelerated
Providing functional, high-purity recombinant proteins—including membrane proteins and nanodiscs—to overcome bottlenecks in drug screening and target validation.
Clones
Validated | Reliable | Comprehensive Collection
Ready-to-use clones for streamlined research and development.
Kits
Complete | Convenient | High Sensitivity
Chromogenic LAL Endotoxin Assay Kit ensures precise, FDA-compliant endotoxin quantification for biosafety testing.
Enzymes
Purified | Stable | Efficient
Powerful Tn5 Transposase for DNA insertion and random library construction.
Aptamers
Highly Specific | Robust | Versatile
Aptamers for key proteins like ACVR1A, Akt, EGFR, and VEGFR.
Adjuvants
Enhancing | Synergistic | Effective
Enhance immune responses with high-purity, potent CpG ODNs.
Laboratory Equipment
Innovative | Reliable | High-Precision
Effortlessly streamline DNA extraction with CB™ Magnetic-Nanoparticle Systems.
Stable Cell Line Generation
Reliable | Scalable | Customizable
Fast proposals, regular updates, and detailed reports; strict quality control, and contamination-free cells; knockout results in 4-6 weeks.
Target-based Drug Discovery Service
Innovative | Comprehensive | Efficient
Target identification, validation, and screening for drug discovery and therapeutic development.
Custom Viral Service
Versatile | High-Yield | Safe
Unbeatable pricing, fully customizable viral packaging services (covering 30,000+ human genes, 200+ mammals, 50+ protein tags).
Custom Antibody Service
Precise | Flexible | Efficient
End-to-end antibody development support, from target to validation, enabling clients to rapidly obtain application-ready antibodies.
Antibody-Drug Conjugation Service
Integrated | Controlled | Translational
Comprehensive solutions covering design, development, and validation to ensure conjugated drugs with consistent quality and clinical potential.
Protein Degrader Service
Efficient | High-Precision | Advanced Therapeutics
Harness the power of protein degraders for precise protein degradation, expanding druggable targets and enhancing therapeutic effectiveness for cutting-edge drug discovery.
Nucleotides Service
Accurate | Flexible | High-Quality
Custom synthesis of oligonucleotides, primers, and probes for gene editing, PCR, and RNA studies.
Custom RNA Service
Custom RNA ServicePrecise | Flexible | GMP-ReadyCustom
RNA design, synthesis, and manufacturing—covering mRNA, saRNA, circRNA, and RNAi. Fast turnaround, rigorous QC, and seamless transition from research to GMP production.
Custom Libraries Construction Service
Comprehensive | High-throughput | Accurate
Custom cDNA, genomic, and mutagenesis libraries for drug discovery, screening, and functional genomics.
Gene Editing Services
Precise | Efficient | Targeted
Gene editing solutions for gene editing, knockouts, knock-ins, and customized genetic modifications. Integrated multi-platform solutions for one-stop CRISPR sgRNA library synthesis and gene screening services
Microbe Genome Editing Service
Precise | Scalable | Customizable
Enhance microbial productivity with advanced genome editing using Rec-mediated recombination and CRISPR/Cas9 technologies.
Biosafety Testing Service
Reliable | Comprehensive | Regulated
Complete biosafety testing solutions for gene therapy, viral vectors, and biologics development.
Plant Genetic Modification Service
Advanced | Sustainable | Tailored
Genetic modification for crop improvement, biotechnology, and plant-based research solutions.
Plant-based Protein Production Service
Efficient | Scalable | Customizable
Plant-based protein expression systems for biopharmaceuticals, enzyme production, and research.
Aptamers Service
Innovative | Fast | Cost-Effective
Revolutionizing drug delivery and diagnostic development with next-generation high-throughput aptamer selection and synthesis technologies.
CGT Biosafety Testing
Comprehensive | Accurate | Regulatory-compliant
Internationally certified evaluation system for biologics, gene therapies, nucleic acid drugs, and vaccines.
Pandemic Detection Solutions
Rapid | Precise | Scalable
Balancing accuracy, accessibility, affordability, and rapid detection to safeguard public health and strengthen global response to infectious diseases.
cGMP Cell Line Development
Reliable | Scalable | Industry-leading
Stable expression over 15 generations with rapid cell line development in just 3 months.
Supports adherent and suspension cell lines, offering MCB, WCB, and PCB establishment.
GMP mRNA Production
Efficient | Scalable | Precise
Scalable mRNA production from milligrams to grams, with personalized process design for sequence optimization, cap selection, and nucleotide modifications, all in one service.
GMP Plasmid Production
High Quality | Scalable | Regulatory-compliant
Our plasmid production services span Non-GMP, GMP-Like, and GMP-Grade levels, with specialized options for linearized plasmids.
GMP Viral Vector Manufacturing
Scalable | High Yield | Quality-driven
Advanced platforms for AAV, adenovirus, lentivirus, and retrovirus production, with strict adherence to GMP guidelines and robust quality control.
AI-Driven Gene Editing and Therapy
Innovative | Precision | Transformative
AI-powered one-click design for customized CRISPR gene editing strategy development.
AI-Antibody Engineering Fusion
Next-Generation | Targeted | Efficient
AI and ML algorithms accelerate antibody screening and predict new structures, unlocking unprecedented possibilities in antibody engineering.
AI-Driven Enzyme Engineering
Smart | Efficient | Tailored
High-throughput enzyme activity testing with proprietary datasets and deep learning models for standardized and precise enzyme engineering design.
AI-Enhanced Small Molecule Screening
Predictive | Efficient | Insightful
Leverage AI to uncover hidden high-potential small molecules, prioritize leads intelligently, and reduce costly trial-and-error in early drug discovery.
AI-Driven Protein Degrader Drug Development
Innovative | Targeted | Accelerated
Use AI-guided design to optimize protein degraders, addressing design complexity and enhancing efficacy while shortening development timelines.
| Cat.No. | Product Name | Price |
|---|
| Cat.No. | Product Name | Price |
|---|
| Cat.No. | Product Name | Price |
|---|
| Cat.No. | Product Name | Price |
|---|
BRCA1 was identified in early 1990’s as tumor suppressor, which is known to play a huge role in sustaining the genome stability in many different tissues. BRCA1 mutations are inheritable and the mutation associated with many cancers, such as breast cancer and ovarian cancer. There's a 50–60% chance of getting breast cancer and 50% risk of developing ovarian cancer for BRCA1mutation carriers. BRCA1 mutant tumors showed a high prevalence in the age group of 30 to 50 years. Although the cause of the early onset is unclear, it is generally believed that hormonal factors play a huge role in inducing BRCA1 related breast tumors. The cause for the tissue specific tumorigenecity in BRCA1 mutation could be linked to estrogen receptorα (ER-α) which is repressed by BRCA1 and mutation in BRCA1 leads to ER-α mediated tumorigenesis as breast/ovarian tissues are under the control of exposure to estrogen that is expressed during menstrual cycle.
BRCA1 combines with other tumor suppressors
Yasmeen et al. has shown that the presence of mutant type BRCA1 down regulated the expression of E-cadherin. The reduction of e-cadherin expression has been a hallmark of the EMT process and is associated with the migration and invasion of breast cancer cells. The key transcription factors Snail 1, Snail 2, ZEB 1, Ets-1, FOXC2 and Twist are acquired during the process of EMT. Snail is a downstream target of many signaling pathways that leads to EMT during cancer and in fact EMT is considered as a rare event without the activation of Snail/Slug. In breast cancer, the expression of slug is up regulated in highly aggressive basal breast tumors in comparison to luminal breast tumors. Also, the over expression of slug is reported to induce basal phenotype in breast cancer harboring luminal phenotype and vice versa which ultimately drives the process of EMT. In breast tumors which are BRCA1 mutated, slug expression is up regulated and in breast tissues which harbor wild type BRCA1 slug expression is down regulated.
Cytoskeletal proteins play a dominant role during EMT in all types of breast cancers. The over expression and under expression of certain cytoskeletal proteins in BRCA1 mutated tumors confers the basal like aggressive behavior of the tumorThe key cytoskeletal proteins that are linked with the process of EMT are β-catenin, vimentin and cytokeratins, where former two molecules are induced and cytokeratins are attenuated during EMT.
Micro RNA’s are 18-25 nucleotides small non coding RNA’s which play a myriad of functions in the normal and cancerous tissues by post transcriptional regulation of critical genes. There are many evidences show that micro RNA can relate breast cancer with BRCA1. Many micro RNA’s such as miR-205, miR-155, miR-99b can be regulated by BRCA1 and also a few micro RNA’s regulating the expression of BRCA1 such as miR-146a, 146b, miR-182, miR-638, miR-17, miR-335, miR-342 is also reported.
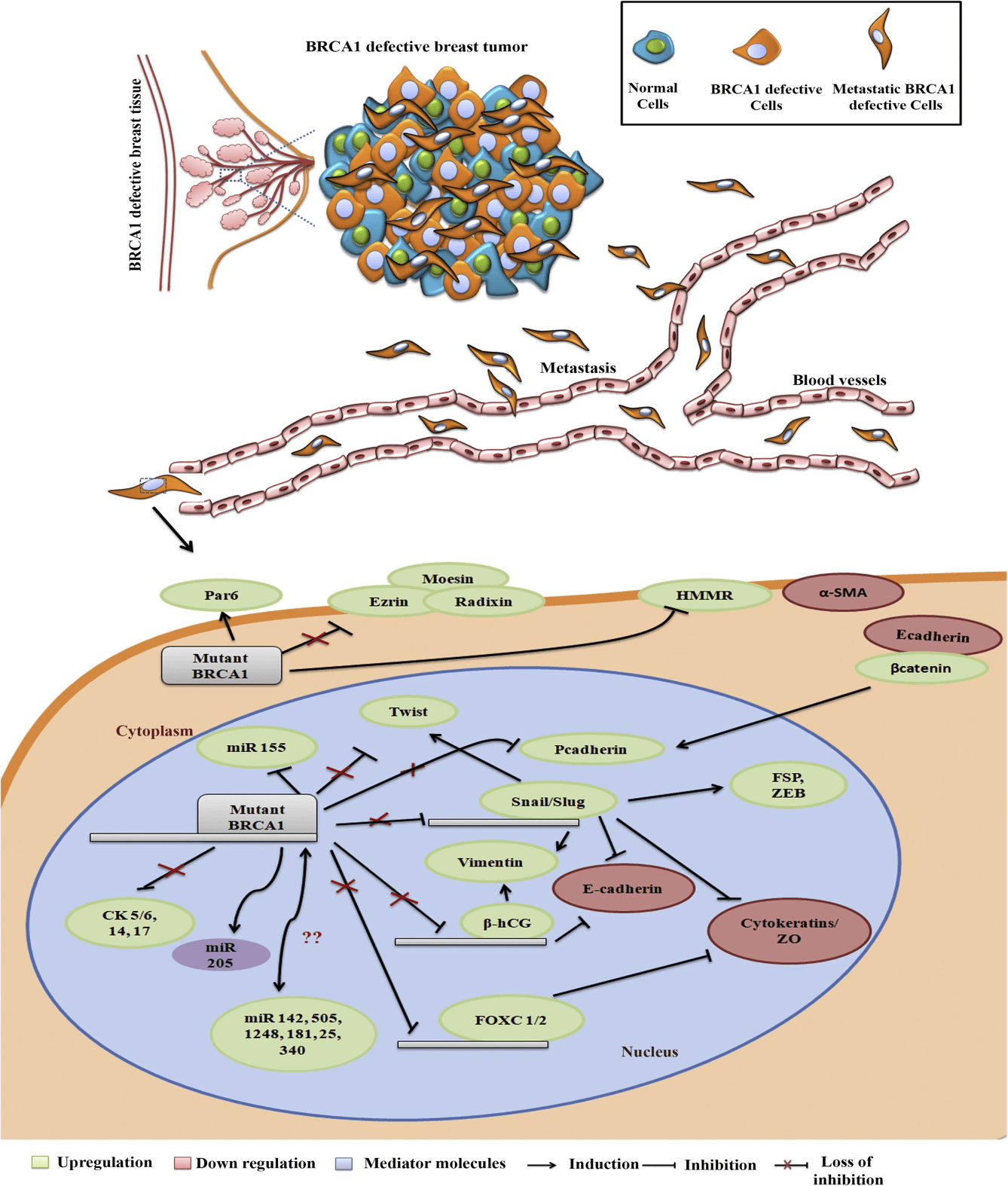 Fig 1. Regulation of EMT molecules in BRCA1 wild type and BRCA1 mutant breast cancer cells. (S.K. Sengodan, et al. 2018).
Fig 1. Regulation of EMT molecules in BRCA1 wild type and BRCA1 mutant breast cancer cells. (S.K. Sengodan, et al. 2018).
BRCA1 play a key role in DNA damage repair
External and internal hazards including reactive oxygen species, ionizing radiation (IR) and many chemicals can induce DNA double strand breaks (DSBs). Various cell cycle checkpoints are activated in response to DSBs or paused replication forks. These checkpoints are considered to coordinate cell cycle progression with DNA repair or induce apoptosis when DNA damage cannot be repaired promptly. Therefore, DNA damage checkpoints are considered important self-defense mechanisms to ensure genome stability. Many proteins including the protein kinase ataxiatelangiectasia mutated (ATM), γH2AX, mediator of DNA damage checkpoint protein 1 (MDC1), Nijmegen breakage syndrome 1 (NBS1), the product of Breast Cancer Susceptibility Gene 1 (BRCA1), and checkpoint kinases 1 and 2 (Chk1 and Chk2) are involved in the ionizing radiation (IR)-induced DNA damage response pathway.
The BRCA1 gene encodes all 1863 amino acid protein, which contains two specific regions, the N-terminal Ring domain that has E3 ligase activity and the tandem C terminal BRCT motifs that specifically recognize phosphoserine/ theronine motifs. Following DNA damage, the key upstream sensor ATM initiates the DNA damage response. ATM is activated and autophosphorylated at several residues including Ser 1981 in response to DNA double stranded breaks. As soon as ATM activation, ATM can phosphorylate a histone H2A variant H2AX at residue Ser 139 at or near the sites of DNA breaks. Hence phosphorylated H2AX has been widely used as a marker of DNA double stranded breaks. This phosphorylated H2AX can bind directly to MDC1 BRCT domains and thus accumulate MDC1 to DNA damage sites. MDC1 can directly or indirectly recruit additional active ATM to the sites of DNA damage. This accumulation of active ATMs at DSBs should result in the further phosphorylation of H2AX at mega-base region surrounding any single DNA break and provide docking sites not only for MDC1, but also other checkpoint proteins at or near the sites of DNA breaks. We believe that this positive feedback mechanism amplifies the DNA damage signals and is especially critical for checkpoint activation following low doses of DNA damage.
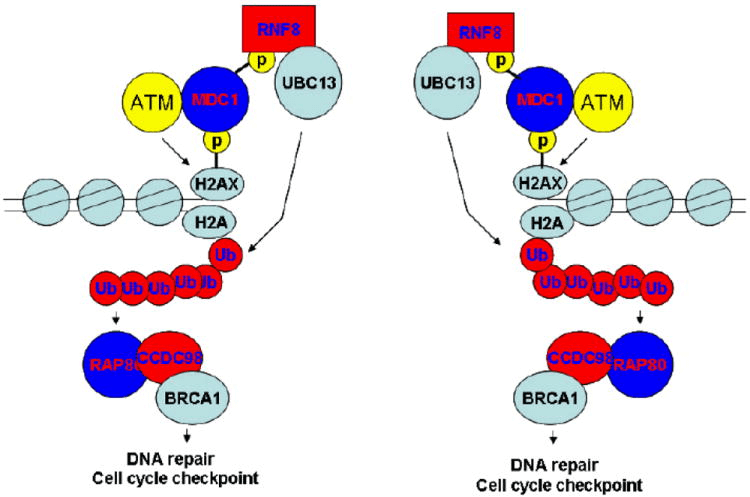 Fig 2. A proposed model of the DNA-damage response pathway (Kim and Chen. 2008).
Fig 2. A proposed model of the DNA-damage response pathway (Kim and Chen. 2008).
References:
Contact us today for a free consultation with the scientific team and discover how Creative Biogene can be a valuable resource and partner for your organization.
Inquiry

Copyright © Creative Biogene. All rights reserved.